About TTSMI (TTS Mapping and Integration)
A triplex target DNA site (TTS), a stretch of DNA sequence composed of poly-purines, is able to selectively form triple-helix (triplex) structure with triplex-forming oligonucleotide (TFO) and influence site-specific modulation of gene expression, site-specific mutagenesis, and modification of genomic DNA as shown in Figure below. For modulation of gene expression, gene regulatory elements can be targeted specifically with TFO to form triplex structure which interferes with either the binding of transcription factors or the formation of the transcription initiation complex, consequently leading to modulation of the gene transcription. For site-specific mutagenesis and modification of genomic DNA, TFO (such as peptide nucleic acids (PNA) and TFO conjugated to psoralen) have been used to induce site-specific DNA damage, thereby stimulating DNA repair system which results in targeted mutagenesis, targeted recombination and sequence-selective manipulation of genomic DNA.
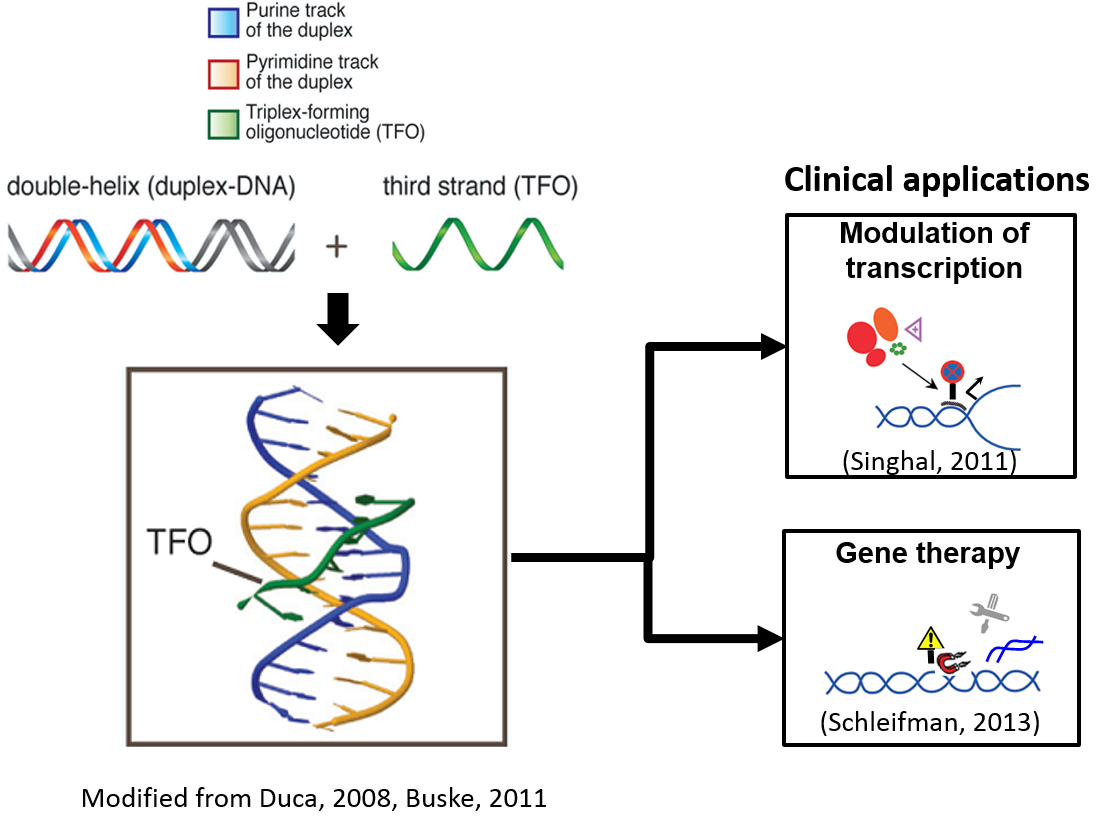
Therefore, co-localization of genomic TTS with gene regulatory signals and functional genome structures suggests the potential impact of TFOs and its functional associability to their TTS targets which in turn could be exploited in antigene strategies for gene therapy of cancers and other genetic diseases. In the field of anti-gene therapy, specific TTS could be targeted by TFO in order to inhibit functions of the regulatory DNA sites (e.g. in gene promoter) and elongation process.
TTSMI (TTS Mapping and Integration) database allows the user to facilitate in (i) searching of the TTS using several search terms including genomic location, gene ID, NCBI RefSeq ID, TTS ID, gene symbol and gene description keywords, (ii) interactive filtering of the TTS co-localized with several other gene regulatory signals, (iii) exploring of dynamic combination of structural and functional annotations of specific TTS, and (iv) viewing of the TTS simultaneously with diverse annotation tracks via UCSC genome browser link.
With the development of the TTSMI database for the human genome, we provide the scientific and industrial communities with new resource to identify unique location of TTS which are integrated with genomic functional annotations and also non-B DNA structures. The user can quickly search and filter the TTS candidates from our largest collection of unique TTS location. In terms of finding therapeutic targets, the specificity of TTS is the most important factor to control off-target effects. Therefore we provided only uniquely located TTS in the genome and simultaneously also provide an off-target report to ensure that users will choose to target a TTS that minimizes off-target effects. We believe that the TTSMI database will facilitate non-bioinformaticians to easily find acceptable TTS candidates for antigene therapy development via triplex technology.
Imprint
Disclaimer: TTSMI Databases and associated information are protected by copyright. This server and its associated data and services are for academic, non-commercial use only. The Bioinformatics Institute /A*STAR has no liability for the use of results, data or information which have been provided through this server. Neither the use for commercial purposes, nor the redistribution of TTSMI database files to third parties, nor the distribution of parts of files or derivative products to any third parties are permitted.
